
_Andrew Cohen
Cohen is an associate professor of electrical and computer engineering in the College of Engineering.
One of the longstanding debates in science that has permeated into the field of stem cell research is the question of nature versus nurture influencing development. Science on stem cells thus far has suggested that their fate is not predestined. But new research from the New York-based Neural Stem Cell Institute and Drexel’s College of Engineering suggests otherwise.
In a recent paper published in Stem Cell Reports, researchers present voluminous video evidence to support their discovery of distinct, intrinsic differences among stem cells found in the brains of mice. Through time-lapse imaging of cells cultured in identical in vitro environments, researchers were able to identify differences in growth, movement and differentiation that exist between stem cells found in two different areas of the brain.
Neural stem cells are some of the most fascinating and unique cells in the body, but very little is known about their unusual behavior. A research group at the Neural Stem Cell Institute in New York studied single neural stem cells from the anterior and posterior embryonic mouse cerebral cortex. They provided the cells with an identical environment in tissue culture, then made time-lapse movies of how they divided and differentiated into cortical cells.
“If the answer to the age-old question was, in fact, ‘nurture’ then these cells would behave the same, given the identical environments,” says Sally Temple, who led the group at the institute. “But if the answer was ‘nature’ then they would behave differently. And this is actually what we found — anterior and posterior stem cells are intrinsically quite different from each other.”
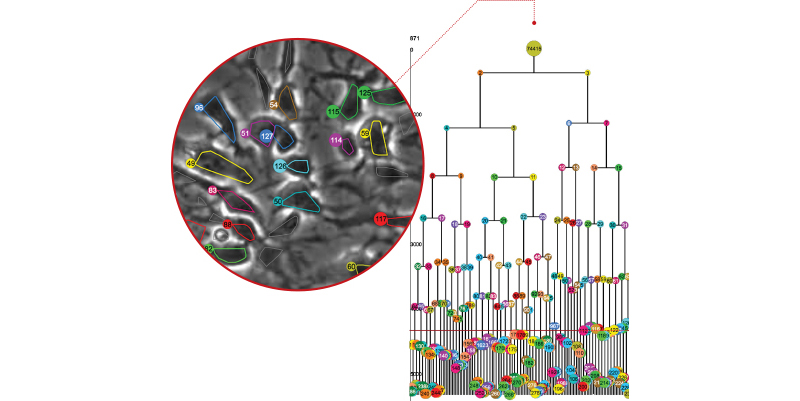
_Family Tree
Drexel’s computational image analysis programs can generate a stem cell lineage tree from a series of time-lapse microscopic images.
This discovery was made possible by a powerful set of biological tracking programs, developed in Drexel’s Computational Image Sequence Analysis Lab led by Andrew Cohen, an associate professor in the College of Engineering. The programs, called LEVER and CloneView, automates the process of analyzing tens of thousands of time-lapse images. They can track the growth, movement and proliferation of cells and use this information to visually enhance the time-lapse movies and produce an entire family tree for each cell.
“Basically, the algorithms look for groups of pixels that have the right appearance and that are in the right place to be the cells we’re tracking,” Cohen says. “From there we use other algorithms to cross-check the results multiple times, until we’re almost 99 percent sure that the things LEVER has identified in all these pictures are, in fact, cells.”
Displayed in CloneView, a time-lapse movie of a single stem cell is overlaid by a uniquely colored outline of its inner boundary. As it grows and eventually divides, new cells are delineated with a different color. Each of these colors corresponds to the cell’s node on the lineage tree, which is displayed in a separate frame next to the movie. This gives researchers a quick way to check the video to make sure LEVER correctly identified the cells and their progeny.
Once checked by human eyes, the images are run through the program again for more precise delineation of the cells — this process is especially arduous when tracking stem cells because they often have long arm-like appendages called processes that can look like a separate cell.
“Human validation is a very important part of our process. We are still the most accurate judge of when cells have divided,” Cohen said. “By running the time-lapse images through the program we get an output video that’s easy to visually verify. As these corrections are made, the program can spot related errors and automatically fix those. The key is to minimize the amount of effort required from the humans.”